This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
|
Click to enlarge phylogenetic trees.
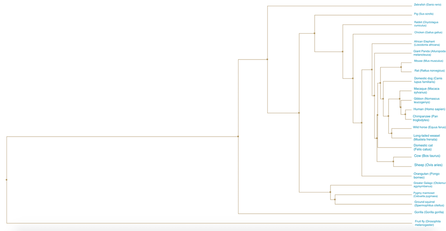
Figure 1. Average distance using BLOSUM62. A phylogenetic tree constructed using average distance and the BLOSUM matrix for the SEMA5A protein.
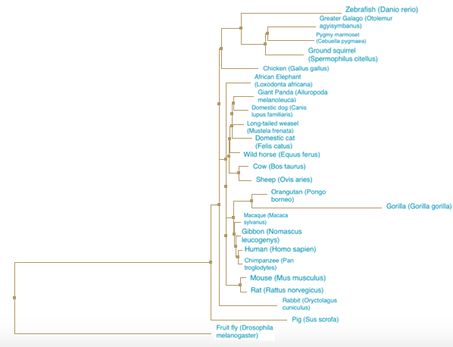
Figure 2. Neighbor joining using BLOSUM62. A phylogenetic tree constructed using neighbor joining and the BLOSUM matrix for the SEMA5A protein.
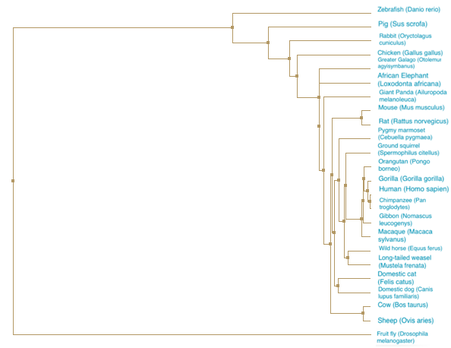
Figure 3. Average distance using percent identity. A phylogenetic tree constructed using average distance and the percent identity score for the SEMA5A protein.
|
What is protein Phylogeny?Protein phylogeny can be used to determine the relatedness of different species, based on their protein sequences, over time. A protein phylogenetic tree shows the evolutionary relationships of different species based on the similarity or differences in the protein sequences. Many methods can be used to determine the similarity between sequences. Only a few methods will be described here.
Generating a phylogenetic treePercent IdentityThe 'percent identity' method looks at the percentage that two sequences are identical to one another. After two sequences have been aligned, the percentage that the same amino acids are found in the same positions in the two sequences, is calculated. Once the similarity scores are calculated, a phylogenetic tree can be generated using several different methods. 'Neighbor Joining' and 'Average Distance' are the methods discussed here [1] [2].
BLOSUM MAtrixThe 'BLOSUM matrix' method is used to calculate the similarity between protein sequences. After two sequences have been aligned, the BLOSUM matrix is used to assign a score to each pair of aligned amino acids, based on the likelihood that these two amino acids would match by random chance. The BLOSUM62 matrix used to assign scores can be found here. The scores at all the sites are summed to give a total score that defines the relatedness of the two sequences. A higher score indicates sequences that are more closely related. Once the similarity scores are calculated, a phylogenetic tree can be generated using several different methods. 'Neighbor Joining' and 'Average Distance' are the methods discussed here [2] [3].
Neighbor JoiningThe 'neighbor joining' method uses the similarity scores calculated by 'BLOSUM' or 'percent identity' to determine the relatedness between species. The amount of change that follows the divergence of the two species is calculated to determine the branch lengths for the phylogenetic tree. This creates a tree with varying branch lengths [2] [4].
Average distanceThe 'average distance' method uses the similarity scores calculated by 'BLOSUM' or 'percent identity' to determine the most closely related species. The most closely related species are joined with equal branch lengths to create a node. The 'average distance' method makes the assumption that both species equally diverged from their common ancestor [4].
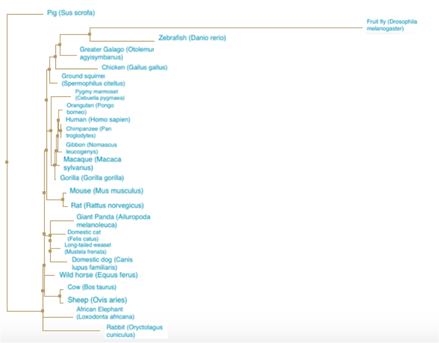
DiscussionThe neighbor joining and average distance phylogenetic trees had relatively similar relationships between between species. In all four trees, the Drosophila melanogaster (fruit fly) is the most distantly related to all other species. The primates and rodents are consistently seen as closely related. All the trees follow what would be expected of a typical species tree, indicating the fidelity of the relationships demonstrated by the trees of the SEMA5A protein.
|
|
Figure 4. Neighbor joining using percent identity. A phylogenetic tree constructed using neighbor joining and the percent identity score for the SEMA5A protein.
|
References
[1] Fassler, J. (2011, July 14). BLAST Help. Retrieved March 7, 2015, from http://www.ncbi.nlm.nih.gov/books/NBK62051/
[2] Professor Ahna Skop's Website: http://genetics564.weebly.com/homology--phylogeny.html
[3] Eddy, S. (n.d.). Where did the BLOSUM62 alignment score matrix come from? Nature Biotechnology, 22, 1035-1036.
[4] Barton, N. (2007, January 1). Phylogenetic Reconstruction. Retrieved March 7, 2015, from http://evolution-textbook.org/content/free/contents/ch27.html#ch27-4-2
[2] Professor Ahna Skop's Website: http://genetics564.weebly.com/homology--phylogeny.html
[3] Eddy, S. (n.d.). Where did the BLOSUM62 alignment score matrix come from? Nature Biotechnology, 22, 1035-1036.
[4] Barton, N. (2007, January 1). Phylogenetic Reconstruction. Retrieved March 7, 2015, from http://evolution-textbook.org/content/free/contents/ch27.html#ch27-4-2