This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
What iS Gene Homology?
A homologous gene is when two species share the same gene as a common ancestor. Gene homology can be used to understand the structure, physiology, or development of different species of organisms [1]. Gene homology can be used to easily research disease, development, and a number of other topics. Model organisms with relatively similar gene sequences, called homologs, can be used for research studies that cannot be conducted on humans. The use of model organisms can accelerate the research and pharmaceutical testing.
SEMA5A Gene Sequences
BLAST Alignment
BLAST, or The Basic Local Alignment Search Tool, looks at the similarity between multiple nucleotide or protein sequences. BLAST compares these sequences to other sequence databases to statistically calculate matches. The results can be used to analyze gene functionality and evolutionary events between sequences [1].
The BLAST results are displayed by [2]:
The BLAST results are displayed by [2]:
- E-value- the probability that the match occurs by random chance
- Percent similarity- the percent of the nucleotide or protein sequence that matched
- Percent identity- the percent of base pairs or amino acids that matched
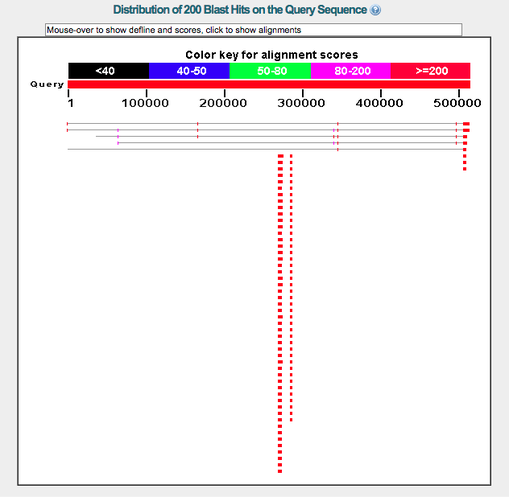
Figure 1. BLAST alignment. This is an example of the distribution of 200 BLAST hits from the SEMA5A gene sequence. The results are color coded, by score, which is indicated by the color key.
Clustal Omega Alignment
Clustal Omega uses multiple sequence alignment to highlight the similarity between sequences so that the conservation of each region can be determined. The results can be used for phylogenetic analysis, which can determine when evolutionary events between sequences might have occurred.
Due to the length of the SEMA5A gene, the gene sequences for this project were unable to be aligned. However, Clustal Omega was used for multiple sequence alignment of the SEMA5A protein. For the results of SEMA5A protein alignment, see the Protein Homology page.
Due to the length of the SEMA5A gene, the gene sequences for this project were unable to be aligned. However, Clustal Omega was used for multiple sequence alignment of the SEMA5A protein. For the results of SEMA5A protein alignment, see the Protein Homology page.
Discussion
The SEMA5A gene was found to be relatively well conserved among vertebrates and some non-vertebrates. These results explain why a chromosomal deletion of the SEMA5A gene, which results in a loss of SEMA5A protein, has drastic phenotypic results, as seen in individuals with Cri du Chat syndrome. It should be noted that the SEMA5A protein is more highly conserved than the SEMA5A gene, which can be seen in the results from the BLAST alignment.
References
Figure 1. Stephen F. Altschul, Thomas L. Madden, Alejandro A. Schäffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of protein database search programs", Nucleic Acids Res. 25:3389-3402.
Figure 2. Sievers F, Wilm A, Dineen DG, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins D
Molecular Systems Biology 7 Article number: 539 doi:10.1038/msb.2011.75
[1] Homology | evolution. (n.d.). Retrieved March 11, 2015, from http://www.britannica.com/EBchecked/topic/270557/homology
[2] Basic Local Alignment Search Tool. (n.d.). Retrieved February 21, 2015, from http://blast.ncbi.nlm.nih.gov/Blast.cgi
[3] Wheeler D, Bhagwat M. BLAST QuickStart: Example-Driven Web-Based BLAST Tutorial. In: Bergman NH, editor. Comparative Genomics: Volumes 1 and 2. Totowa (NJ): Humana Press; 2007. Chapter 9. Available from: http://www.ncbi.nlm.nih.gov/books/NBK1734/
Figure 2. Sievers F, Wilm A, Dineen DG, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins D
Molecular Systems Biology 7 Article number: 539 doi:10.1038/msb.2011.75
[1] Homology | evolution. (n.d.). Retrieved March 11, 2015, from http://www.britannica.com/EBchecked/topic/270557/homology
[2] Basic Local Alignment Search Tool. (n.d.). Retrieved February 21, 2015, from http://blast.ncbi.nlm.nih.gov/Blast.cgi
[3] Wheeler D, Bhagwat M. BLAST QuickStart: Example-Driven Web-Based BLAST Tutorial. In: Bergman NH, editor. Comparative Genomics: Volumes 1 and 2. Totowa (NJ): Humana Press; 2007. Chapter 9. Available from: http://www.ncbi.nlm.nih.gov/books/NBK1734/