This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
Introduction
Cri du Chat syndrome is a chromosomal deletion disease that, while rare, can be devastating for an affected family. The syndrome is caused by a homozygous deletion in the short arm of the fifth chromosome [1]. Mild cases of Cri du Chat syndrome are manageable with therapy, but more severe chromosomal deletions can result in critical medical conditions or organ defects, which are often fatal. Because one of the genetic causes of Cri du Chat syndrome is the deletion of SEMA5A, the primary goal of the specific aims outlined here is to determine the proteomic changes that contribute to abnormal vascularization, which is potentially involved in fatal organ defects in individuals with Cri du Chat. This research could potentially discover patterns that lead to abnormal vascular development in Cri du Chat, Autism, Parkinson’s disease, and mental retardation patients.
Previous studies have shown that SEMA5A deletion can lead to abnormal brain development, but there have been few studies looking at SEMA5A involvement in other organ systems [2]. A recent study showed that SEMA5A knockout mice (Mus musculus) die early because of SEMA5A involvement in vascular advancement in embryonic development [3]. SEMA5A mutant mice exhibit abnormal vasculature localization, suggesting that SEMA5A is involved in vascular positioning. Although there have been many studies on abnormal brain development associated with SEMA5A deletion, the functions of SEMA5A in vascular development are still unknown. The goals of this study are to research the significance of SEMA5A in vascular patterning during embryonic development. This will be studied using proteomic and bioinformatic tools, which will be applied to the human and fruit fly proteome. I hypothesized that the SEMA5A protein is necessary for proper development in organisms with circulatory systems.
Previous studies have shown that SEMA5A deletion can lead to abnormal brain development, but there have been few studies looking at SEMA5A involvement in other organ systems [2]. A recent study showed that SEMA5A knockout mice (Mus musculus) die early because of SEMA5A involvement in vascular advancement in embryonic development [3]. SEMA5A mutant mice exhibit abnormal vasculature localization, suggesting that SEMA5A is involved in vascular positioning. Although there have been many studies on abnormal brain development associated with SEMA5A deletion, the functions of SEMA5A in vascular development are still unknown. The goals of this study are to research the significance of SEMA5A in vascular patterning during embryonic development. This will be studied using proteomic and bioinformatic tools, which will be applied to the human and fruit fly proteome. I hypothesized that the SEMA5A protein is necessary for proper development in organisms with circulatory systems.
Aim 1
The purpose of the first aim was to determine predicted and conserved post-translational modifications in SEMA5A protein between different vascular organisms. Identification of important phosphorylation sites in SEMA5A will help determine the importance of SEMA5A in vascular patterning because several signaling pathways depend on post-translational modifications for activity.
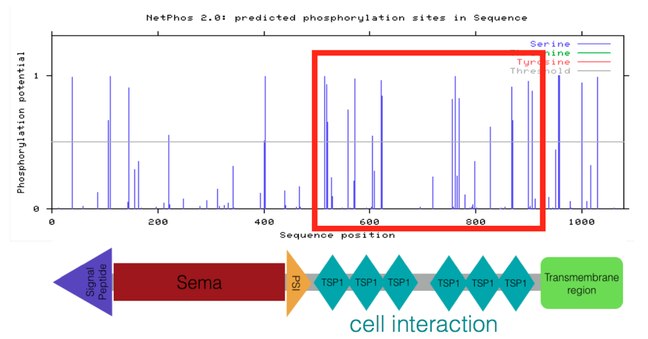
A NetPhos 2.0 analysis was performed to predict phosphorylation sites in human SEMA5A protein. NetPhos 2.0 predicts phosphorylation sites in a protein sequence based on phosphorylatability of an amino acid. Phosphorylatable amino acids include: serine, tyrosine, and threonine. Each site is assigned a score between zero and one (with one being high likelihood that the predicted site is phosphorylated) [4]. This aim focused specifically on the phosphorylation of serine sites in the TSP1 protein domain region because I hypothesized this region (which is involved in regulation of cell interactions in vertebrates) is important for vascular patterning during development. Below are the phosphorylation prediction score results for the phosphorylation of serine sites from NetPhos 2.0:
A NetPhos 2.0 analysis was performed to predict phosphorylation sites in human SEMA5A protein. NetPhos 2.0 predicts phosphorylation sites in a protein sequence based on phosphorylatability of an amino acid. Phosphorylatable amino acids include: serine, tyrosine, and threonine. Each site is assigned a score between zero and one (with one being high likelihood that the predicted site is phosphorylated) [4]. This aim focused specifically on the phosphorylation of serine sites in the TSP1 protein domain region because I hypothesized this region (which is involved in regulation of cell interactions in vertebrates) is important for vascular patterning during development. Below are the phosphorylation prediction score results for the phosphorylation of serine sites from NetPhos 2.0:
Figure 1. Potential phosphorylation scores for serines in human SEMA5A.
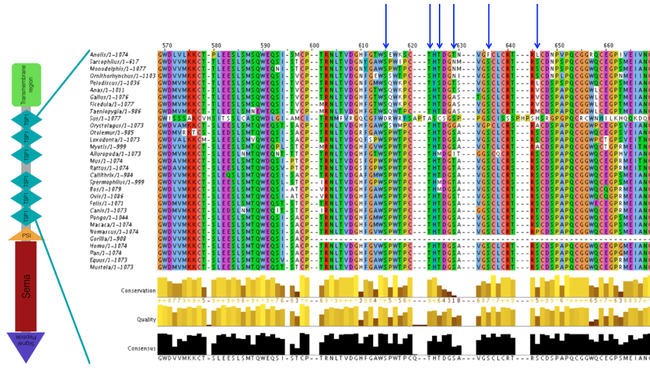
To determine the conserved post-translational modifications between different vascular organisms, I align protein sequences in Clustal Omega. Clustal Omega uses multiple sequence alignment to highlight the similarity between sequences so that the conservation of each region can be determined. Below are the Clustal Omega alignment results (from the TSP1 domain region), with six highly conserved serine and threonine sites that have some of the highest prediction scores (highlighted by the blue arrows):
Figure 2. Clustal Omega alignment results from the TSP1 protein domain region. The blue arrows indicate highly conserved serine and threonine sites that could be functionally necessary for vascular patterning in early development.
I would suggest the use of mass spectrometry to validate, in vivo, the six predicted phosphorylated sites highlighted in this aim.
Aim 2
The purpose of this second aim was to determine unique protein interactions of SEMA5A that contribute to proper vascular development. Identification of unique protein interactions will help determine the important SEMA5A protein interactions involved in vascular patterning.
To isolate protein complexes, I would suggest co-immunoprecipitation (identifies protein-protein interactions by using target protein-specific antibodies to capture proteins that are bound to a specific target protein [5]) or tandem affinity purification tagging (identifies protein-protein interaction by purifying protein complexes [6]) on fruit flies (Drosophila melanogaster) with the homologous SEMA5A protein. Tandem mass spectrometry could then be utilized to identify subsequent proteins.
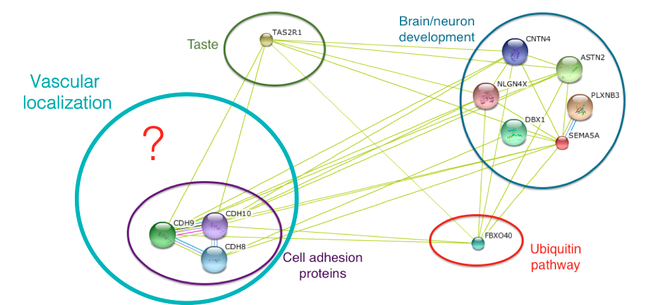
The identified protein interactions will then be compared to the GO analysis already performed for human SEMA5A. The new protein interactions can then be added to String Database to make protein interaction networks that visually represent the unique protein-protein interactions. It would be expected that more cell adhesion protein interactions would be identified because cell adhesion proteins have been verified as important proteins for vascularization. Below are the hypothetical results from this aim:
To isolate protein complexes, I would suggest co-immunoprecipitation (identifies protein-protein interactions by using target protein-specific antibodies to capture proteins that are bound to a specific target protein [5]) or tandem affinity purification tagging (identifies protein-protein interaction by purifying protein complexes [6]) on fruit flies (Drosophila melanogaster) with the homologous SEMA5A protein. Tandem mass spectrometry could then be utilized to identify subsequent proteins.
The identified protein interactions will then be compared to the GO analysis already performed for human SEMA5A. The new protein interactions can then be added to String Database to make protein interaction networks that visually represent the unique protein-protein interactions. It would be expected that more cell adhesion protein interactions would be identified because cell adhesion proteins have been verified as important proteins for vascularization. Below are the hypothetical results from this aim:
Figure 3. String Database results showing protein-protein interactions using the human SEMA5A protein. Identification of unique protein interactions that are involved in vascular patterning, such as more cell adhesion proteins, would be expected.
The simple vasculature and sequenced genome of Drosophila melanogaster make it an ideal organism for comparison to an organism with complex vasculature, such as Homo sapiens. New cell adhesion proteins that are identified in both Drosophila melanogaster and Homo sapiens could be necessary for proper vascular patterning in vascular organisms.
Aim 3
The purpose of this third aim is to identify conserved phosphorylation sites in the Thrombospondin type 1 repeats domain region (a region potentially involved in vascularization) in complex vascular organisms, humans (Homo sapiens), and simple vascular organisms, fruit flies (Drosophila melanogaster). Identification of conserved phosphorylation sites in SEMA5A will help determine the importance of SEMA5A in vascular patterning.
A NetPhos 2.0 analysis was performed to predict phosphorylation sites in human SEMA5A protein. NetPhos 2.0 predicts phosphorylation sites in a protein sequence based on phosphorylatability of an amino acid. Phosphorylatable amino acids include: serine, tyrosine, and threonine. Each site is assigned a score between zero and one (with one being high likelihood that the predicted site is phosphorylated) [4]. This aim focused specifically on the phosphorylation of serine sites in the TSP1 protein domain region because I hypothesized this region (which is involved in regulation of cell interactions in vertebrates) is important for vascular patterning during development. The results for this can be seen above in Figure 1 of Aim 1.
To determine the conserved post-translational modifications between complex vascular organisms, humans (Homo sapiens), and simple vascular organisms, fruit flies (Drosophila melanogaster), I align protein sequences in Clustal Omega. Clustal Omega uses multiple sequence alignment to highlight the similarity between sequences so that the conservation of each region can be determined. The red arrows highlight seven specific amino acids that are predicted to be phosphorylated and are highly conserved between both humans and fruit flies. Below are the results from the Clustal Omega alignment.
A NetPhos 2.0 analysis was performed to predict phosphorylation sites in human SEMA5A protein. NetPhos 2.0 predicts phosphorylation sites in a protein sequence based on phosphorylatability of an amino acid. Phosphorylatable amino acids include: serine, tyrosine, and threonine. Each site is assigned a score between zero and one (with one being high likelihood that the predicted site is phosphorylated) [4]. This aim focused specifically on the phosphorylation of serine sites in the TSP1 protein domain region because I hypothesized this region (which is involved in regulation of cell interactions in vertebrates) is important for vascular patterning during development. The results for this can be seen above in Figure 1 of Aim 1.
To determine the conserved post-translational modifications between complex vascular organisms, humans (Homo sapiens), and simple vascular organisms, fruit flies (Drosophila melanogaster), I align protein sequences in Clustal Omega. Clustal Omega uses multiple sequence alignment to highlight the similarity between sequences so that the conservation of each region can be determined. The red arrows highlight seven specific amino acids that are predicted to be phosphorylated and are highly conserved between both humans and fruit flies. Below are the results from the Clustal Omega alignment.
Figure 4. Clustal Omega alignment results for humans and fruit flies. The red arrows indicate highly conserved amino acids that are predicted to be phosphorylated. These phosphorylated amino acids could be necessary for proper vascular patterning in vascular organisms.
I would suggest the use of the CRISPR/Cas9 system to substitute the highly conserved serine and threonine sites (that are highlighted by the red arrows) with alanine, an amino acid that cannot be phosphorylated. If these phosphorylated serine and threonine sites are necessary for proper vascular patterning, the alanine-mutated fruit flies will not show proper vascularization in development. Mutation of these specific amino acids would help determine the phosphorylation sites that function in vascular patterning in vascular organisms.
Future Directions
If the SEMA5A gene is found to be involved in localization signal pathways and is necessary for proper vascular positioning, gene therapy could potentially be used to rescue abnormal vascular development in Cri du Chat, Autism, Parkinson's, and mental retardation patients. Additionally, drug screens for compounds that rescue vascular phenotypes found in SEMA5A mutants would be beneficial for future treatments.
If the SEMA5A gene is found to not be involved in localization signal pathways and is necessary for proper vascular positioning, exploring more genes and proteins that are necessary for vascularization would be the next step. The results from the second aim could potentially be a place to start in this research, especially if new cell adhesion proteins are discovered.
This research could potentially discover patterns that lead to abnormal vascular development in Cri du Chat, Autism, Parkinson’s disease, and mental retardation patients. Ultimately, this research could lead to new strategies for prenatal treatment of abnormal vascularization.
If the SEMA5A gene is found to not be involved in localization signal pathways and is necessary for proper vascular positioning, exploring more genes and proteins that are necessary for vascularization would be the next step. The results from the second aim could potentially be a place to start in this research, especially if new cell adhesion proteins are discovered.
This research could potentially discover patterns that lead to abnormal vascular development in Cri du Chat, Autism, Parkinson’s disease, and mental retardation patients. Ultimately, this research could lead to new strategies for prenatal treatment of abnormal vascularization.
Cri du Chat Syndrome Awareness
Youtube 1.20.2015. Cri Du Chat Syndrome Awareness. Retrieved from here.
| ||||||||
References
Youtube video: http://youtu.be/F_dW3voR9ZE
[1] Cerruti Mainardi, P. (2006). Cri du Chat syndrome. Orphanet Journal of Rare Diseases, 1, 33. doi:10.1186/1750-1172-1-33.
[2] Genetic Science Learning Center (2014, June 22) Cri-du-Chat Syndrome. Learn.Genetics. Retrieved January 28, 2015, from http://learn.genetics.utah.edu/content/disorders/chromosomal/cdc/
[3] Fiore, R., Rahim, B., Christoffels, V., Moorman, A., & Puschel, A. (2005). Inactivation of the Sema5a Gene Results in Embryonic Lethality and Defective Remodeling of the Cranial Vascular System. Molecular and Cellular Biology, 25(6), 2310-2319, from http://mcb.asm.org/content/25/6/2310.full
[4] Sequence- and structure-based prediction of eukaryotic protein phosphorylation sites. Blom, N., Gammeltoft, S., and Brunak, S. Journal of Molecular Biology: 294(5): 1351-1362, 1999.
[5] Co-immunoprecipitation (Co-IP). (n.d.). Retrieved May 4, 2015, from https://www.lifetechnologies.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/co-immunoprecipitation-co-ip.html
[6] Puig, O. (2001). The Tandem Affinity Purification (TAP) Method: A General Procedure Of Protein Complex Purification. Methods, 218-229.
[1] Cerruti Mainardi, P. (2006). Cri du Chat syndrome. Orphanet Journal of Rare Diseases, 1, 33. doi:10.1186/1750-1172-1-33.
[2] Genetic Science Learning Center (2014, June 22) Cri-du-Chat Syndrome. Learn.Genetics. Retrieved January 28, 2015, from http://learn.genetics.utah.edu/content/disorders/chromosomal/cdc/
[3] Fiore, R., Rahim, B., Christoffels, V., Moorman, A., & Puschel, A. (2005). Inactivation of the Sema5a Gene Results in Embryonic Lethality and Defective Remodeling of the Cranial Vascular System. Molecular and Cellular Biology, 25(6), 2310-2319, from http://mcb.asm.org/content/25/6/2310.full
[4] Sequence- and structure-based prediction of eukaryotic protein phosphorylation sites. Blom, N., Gammeltoft, S., and Brunak, S. Journal of Molecular Biology: 294(5): 1351-1362, 1999.
[5] Co-immunoprecipitation (Co-IP). (n.d.). Retrieved May 4, 2015, from https://www.lifetechnologies.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/co-immunoprecipitation-co-ip.html
[6] Puig, O. (2001). The Tandem Affinity Purification (TAP) Method: A General Procedure Of Protein Complex Purification. Methods, 218-229.