This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
What is a Protein Domain?
A domain is a conserved structural and/or functional unit of a protein. Domains are typically involved in a specific function or interaction that adds to the overall capacity of a protein. Because similar domains can be found in proteins with a different function, domains have been determined to have variation in regards to biological contexts. Protein domains are used to determine a particular protein's function and the conservation of the protein over time [1].
SEMA5A Domain Structure
The SEMA5A protein contains five domains (with some variation, depending on the source): one signal peptide, one sema, one PS1, six thrombospondin type 1 repeats, and one transmembrane region [2] [3].
Figure 1. The SEMA5A domains from SMART. The pink boxes indicate regions of low sequence complexity.
Figure 2. The SEMA5A domains from Pfam. The red boxes represent the five thrombospondin type 1 repeats.
Sema5a Domain ConservatioN
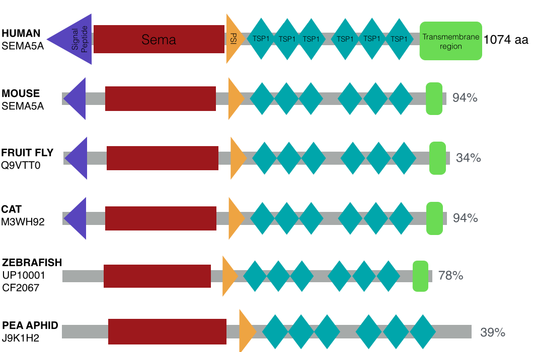
SMART and Pfam were used to determine the protein domains in SEMA5A homologs between a variety of model organisms. The domain alignment shown below was generated based off data in SMART. The depiction also includes the percent identity between the model organism and the human domains. The sema, PS1, and thrombospondin type 1 repeats domains are highly conserved in vertebrates and invertebrates.
Figure 3. The domains in SEMA5A homologs between a variety of model organisms.
Sema5a Domain Functions
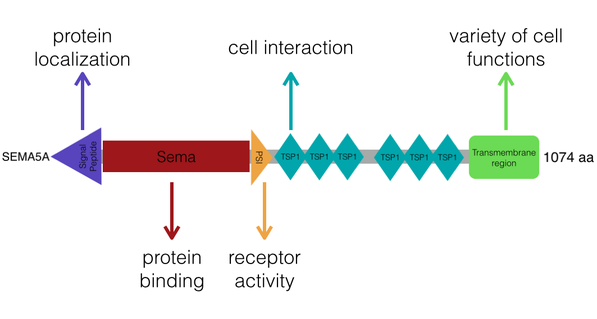
Figure 4. A depiction of the SEMA5A domains and their specific functions, based on the domains determined by SMART.
- Signal Peptide- directs the protein for localization
- Sema- protein binding, specifically during axon guidance
- PSI- receptor activity in multicellular organismal development
- TSPI (Thrombospondin type 1 repeats)- regulators of cell interactions in vertebrates
- Transmembrane region- involved in variety of cellular functions
What is a motif?
A motif is a small unit of a domain or a short region that is assumed to have biological function and is highly conserved. Motifs can be molecular sequences (amino acid or nucleotide sequences) or they can be structural units (secondary protein structures). The study of motifs are important for complete understanding of gene regulation [4] [5].
SEMA5A Protein Motifs
MEME identified three protein motifs in the human SEMA5A protein, all of which were conserved in the all the protein homologs. The results from the MEME analysis can be viewed here. The 'DREME-DNA only' analysis could not be performed due to the length of the gene sequence.
Figure 5. Human SEMA5A domain motif 1. Begins at amino acid residue 990 and ends at residue 1039.
Figure 6. Human SEMA5A domain motif 2. Begins at amino acid residue 224 and ends at residue 273.
Figure 7. Human SEMA5A domain motif 3. Begins at amino acid residue 705 and ends at residue 754.
Discussion
A protein's function can be investigated by protein domains and motifs. SEMA5A contains one signal peptide, one sema, one PS1, six thrombospondin type 1 repeats, and one transmembrane region domains.
The sema, PS1, and thrombospondin type 1 repeats domains are highly conserved in vertebrates and invertebrates. The signal peptide and transmembrane region domains are variable in their evolutionary conservation. The involvement of the sema domain in protein binding, specifically during axon guidance, explains its evolutionary conservation and its importance for proper neuronal development.
There were three motifs found between the SEMA5A protein homologs. Motif 1 is positioned in the transmembrane region domain, while motif 2 is positioned upstream in the sema domain. The positioning of motif 1 supports the information provided by the SMART domains, which indicates the existence of a transmembrane region domain. Motif 3 is positioned at the end of a thrombospondin type 1 repeat domain and the start of a low sequence complexity region.
The sema, PS1, and thrombospondin type 1 repeats domains are highly conserved in vertebrates and invertebrates. The signal peptide and transmembrane region domains are variable in their evolutionary conservation. The involvement of the sema domain in protein binding, specifically during axon guidance, explains its evolutionary conservation and its importance for proper neuronal development.
There were three motifs found between the SEMA5A protein homologs. Motif 1 is positioned in the transmembrane region domain, while motif 2 is positioned upstream in the sema domain. The positioning of motif 1 supports the information provided by the SMART domains, which indicates the existence of a transmembrane region domain. Motif 3 is positioned at the end of a thrombospondin type 1 repeat domain and the start of a low sequence complexity region.
References
[1] Train online: What are protein domains? (n.d.). Retrieved March 25, 2015, from http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains
[2] Select your default SMART mode. (n.d.). Retrieved March 25, 2015, from http://smart.embl-heidelberg.de/
[3] R.D. Finn, A. Bateman, J. Clements, P. Coggill, R.Y. Eberhardt, S.R. Eddy, A. Heger, K. Hetherington, L. Holm, J. Mistry, E.L.L. Sonnhammer, J. Tate, M. PuntaNucleic Acids Research (2014) Database Issue 42:D222-D230, from http://pfam.xfam.org/
[4] Motif - Glossary Entry. (n.d.). Retrieved March 25, 2015, from http://ghr.nlm.nih.gov/glossary=motif
[5] Tak-Ming Chan; Kwong-Sak Leung; Kin-Hong Lee; Lio', P., "Generic spaced DNA motif discovery using Genetic Algorithm," Evolutionary Computation (CEC), 2010 IEEE Congress on , vol., no., pp.1,8, 18-23 July 2010, from http://ieeexplore.ieee.org/xpl/login.jsp?tp=&arnumber=5585924&tag=1&url=http%3A%2F%2Fieeexplore.ieee.org%2Fxpls%2Fabs_all.jsp%3Farnumber%3D5585924%26tag%3D1
[2] Select your default SMART mode. (n.d.). Retrieved March 25, 2015, from http://smart.embl-heidelberg.de/
[3] R.D. Finn, A. Bateman, J. Clements, P. Coggill, R.Y. Eberhardt, S.R. Eddy, A. Heger, K. Hetherington, L. Holm, J. Mistry, E.L.L. Sonnhammer, J. Tate, M. PuntaNucleic Acids Research (2014) Database Issue 42:D222-D230, from http://pfam.xfam.org/
[4] Motif - Glossary Entry. (n.d.). Retrieved March 25, 2015, from http://ghr.nlm.nih.gov/glossary=motif
[5] Tak-Ming Chan; Kwong-Sak Leung; Kin-Hong Lee; Lio', P., "Generic spaced DNA motif discovery using Genetic Algorithm," Evolutionary Computation (CEC), 2010 IEEE Congress on , vol., no., pp.1,8, 18-23 July 2010, from http://ieeexplore.ieee.org/xpl/login.jsp?tp=&arnumber=5585924&tag=1&url=http%3A%2F%2Fieeexplore.ieee.org%2Fxpls%2Fabs_all.jsp%3Farnumber%3D5585924%26tag%3D1