This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
What are protein modifications?
Protein modification occurs when a compound is attached to a protein either during or after translation. Post-translational modifications are involved in several cellular processes, like protein degradation, protein-protein interactions, gene expression regulation, cellular differentiation, or signaling processes. The most well studied post-translational modification is phosphorylation because it is necessary for numerous cellular functions and signaling pathways. Approximately one third of mammalian protein is phosphorylated. Research of post-translational modifications is necessary to better understand the molecular functions of proteins [1].
SEMA5A Post-translational Modifications
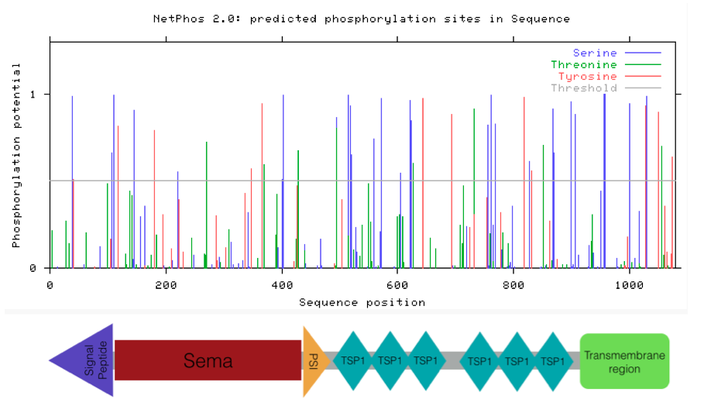
NetPhos 2.0 predicts phosphorylation sites in a protein sequence based on phosphorylatability of an amino acid. Phosphorylatable amino acids include: serine, tyrosine, and threonine. Each site is assigned a score between zero and one (with one being high likelihood that the predicted site is phosphorylated) [2]. There are 29 serine, 12 tyrosine, and 8 threonine predicted phosphorylated sites on the SEMA5A protein.
Figure 1. Predicted phosphorylation sites in human SEMA5A protein.
Discussion
This data suggest that there are many predicted phosphorylation sites in the human SEMA5A protein. There are numerous sites in the TSP1 and transmembrane region domains that have high likelihood of being phosphorylated. More research will need to be conducted to determine the phosphorylation sites that are necessary for proper development. Results from research regarding highly conserved phosphorylation sites that could be involved in vascularization patterning can be found on the Conclusions page.
References
[1] Post Translational Modifications (PTMs). (n.d.). Retrieved April 30, 2015, from http://keck.med.yale.edu/proteomics/technologies/posttransmodifications/index.aspx
[2] Sequence- and structure-based prediction of eukaryotic protein phosphorylation sites. Blom, N., Gammeltoft, S., and Brunak, S. Journal of Molecular Biology: 294(5): 1351-1362, 1999.
[2] Sequence- and structure-based prediction of eukaryotic protein phosphorylation sites. Blom, N., Gammeltoft, S., and Brunak, S. Journal of Molecular Biology: 294(5): 1351-1362, 1999.
Michaela Cline [email protected] Updated: 4.29.2015 Genetics 564